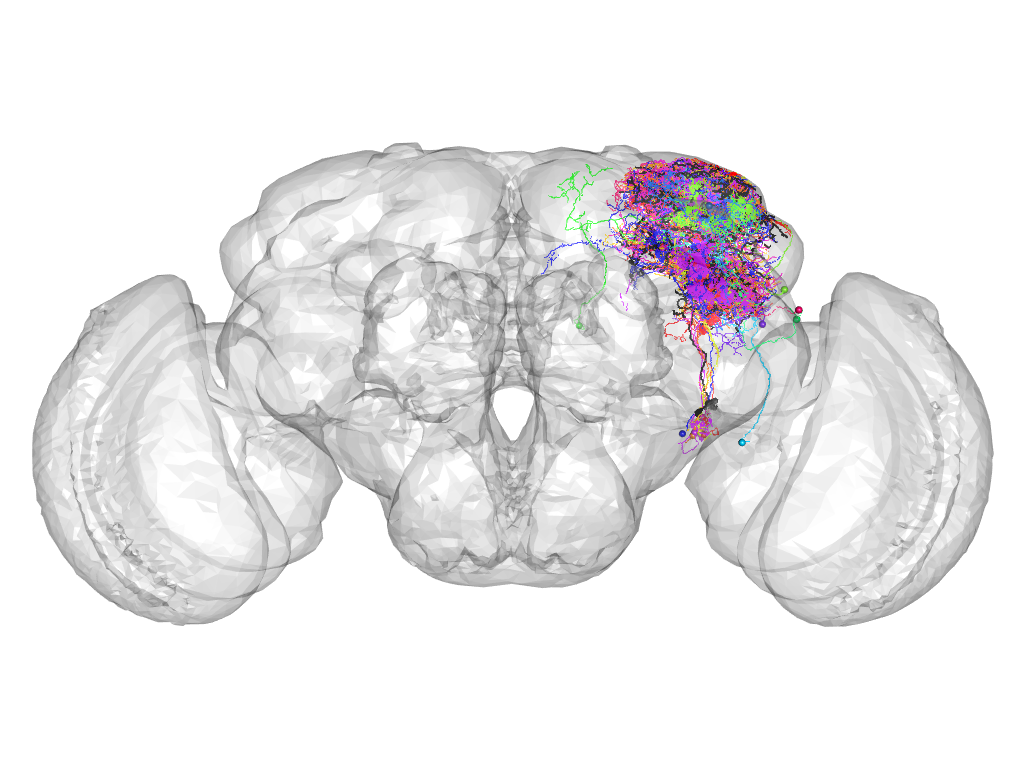
Cluster 194
Part of supercluster XVII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-M-200035), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 136 | TH-M-000013 | XVII | 0.59 |
| 461 | Cha-F-900029 | XVII | 0.49 |
| 268 | Cha-F-200340 | X | 0.39 |
| 440 | VGlut-F-600716 | XVII | 0.32 |
| 984 | VGlut-F-200146 | XVII | 0.31 |
Cluster composition
This cluster has 16 neurons. The exemplar of this cluster (TH-M-200035) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-M-200035 | 3207 | THMARCM-328M_seg1 | TH-Gal4 | M | 141363.12 | 1.00 |
| TH-M-000042 | 3062 | THMARCM-627M-X2_seg2 | TH-Gal4 | M | 100289.32 | 0.72 |
| TH-M-200033 | 3205 | THMARCM-322M_seg1 | TH-Gal4 | M | 108407.08 | 0.77 |
| TH-M-200079 | 3280 | THMARCM-474M_seg2 | TH-Gal4 | M | 103129.37 | 0.75 |
| TH-M-300048 | 3282 | THMARCM-477M_seg1 | TH-Gal4 | M | 90359.21 | 0.71 |
| Cha-F-100357 | 3913 | ChaMARCM-F001572_seg001 | Cha-Gal4 | F | 52616.72 | 0.48 |
| Gad1-F-200151 | 4118 | GadMARCM-F000704_seg001 | Gad1-Gal4 | F | 55221.17 | 0.44 |
| Cha-F-000350 | 4372 | ChaMARCM-F001428_seg001 | Cha-Gal4 | F | 32578.35 | 0.44 |
| Cha-F-100103 | 5512 | ChaMARCM-F000827_seg002 | Cha-Gal4 | F | 45128.33 | 0.53 |
| Cha-F-000152 | 5561 | ChaMARCM-F000861_seg001 | Cha-Gal4 | F | 33900.35 | 0.41 |
| Cha-F-700102 | 7559 | ChaMARCM-F000409_seg001 | Cha-Gal4 | F | 63348.41 | 0.50 |
| Gad1-F-500004 | 10724 | GadMARCM-F000052_seg001 | Gad1-Gal4 | F | 105103.83 | 0.72 |
| VGlut-F-600223 | 11825 | DvGlutMARCM-F002387_seg002 | VGlut-Gal4 | F | 40557.47 | 0.46 |
| TH-F-300052 | 13528 | THMARCM-411F_seg1 | TH-Gal4 | F | 107526.61 | 0.76 |
| TH-F-300067 | 13566 | THMARCM-470F_seg2 | TH-Gal4 | F | 106589.57 | 0.77 |
| VGlut-F-200260 | 14960 | DvGlutMARCM-F1298_seg1 | VGlut-Gal4 | F | 44782.79 | 0.51 |