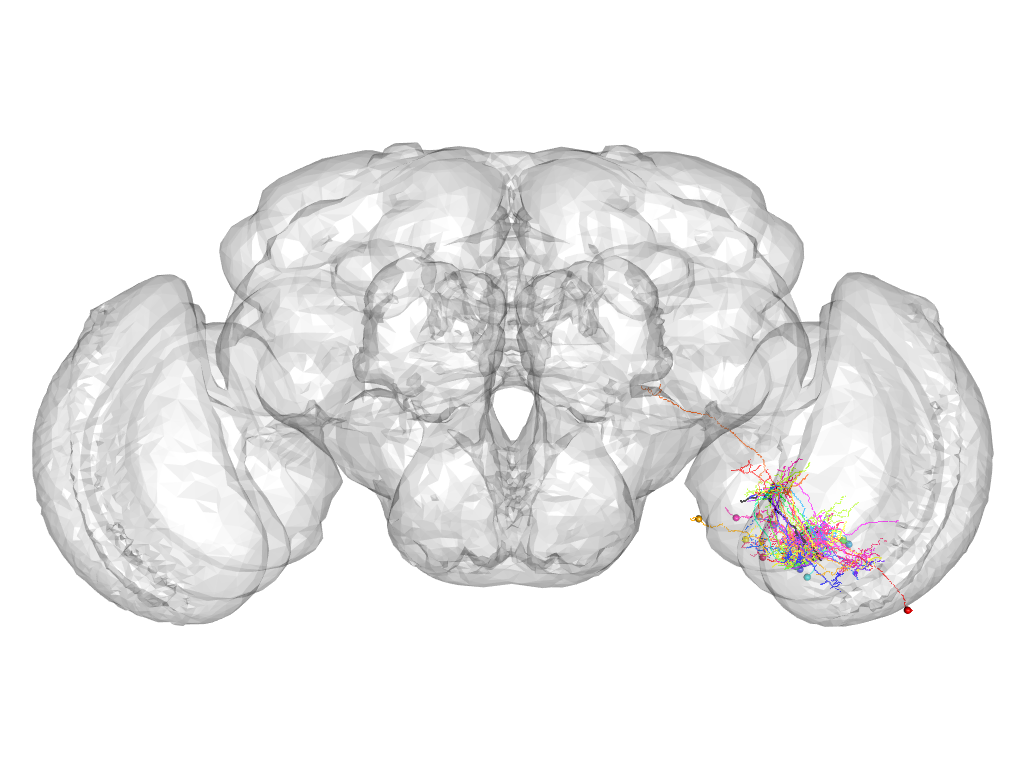
Cluster 1035
Part of supercluster V
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-500088), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 899 | VGlut-F-100044 | V | 0.72 |
| 935 | VGlut-F-100111 | V | 0.65 |
| 133 | fru-M-700043 | I | 0.55 |
| 712 | VGlut-F-900055 | V | 0.53 |
| 316 | fru-F-400204 | IV | 0.50 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (VGlut-F-500088) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-500088 | 16059 | DvGlutMARCM-F446-X2_seg1 | VGlut-Gal4 | F | 11400.62 | 1.00 |
| fru-M-400061 | 1412 | FruMARCM-M000925_seg001 | fru-Gal4 | M | 8437.03 | 0.62 |
| fru-F-200069 | 7383 | FruMARCM-F001027_seg001 | fru-Gal4 | F | 5919.28 | 0.36 |
| VGlut-F-300537 | 7974 | DvGlutMARCM-F003802_seg001 | VGlut-Gal4 | F | 7261.68 | 0.54 |
| VGlut-F-800241 | 7986 | DvGlutMARCM-F003811_seg001 | VGlut-Gal4 | F | 6067.27 | 0.44 |
| VGlut-F-700296 | 8539 | DvGlutMARCM-F003115_seg001 | VGlut-Gal4 | F | 6985.98 | 0.49 |
| VGlut-F-500579 | 8565 | DvGlutMARCM-F003132_seg001 | VGlut-Gal4 | F | 7544.95 | 0.65 |
| VGlut-F-500695 | 8750 | DvGlutMARCM-F003577_seg001 | VGlut-Gal4 | F | 8182.49 | 0.71 |
| VGlut-F-800166 | 9069 | DvGlutMARCM-F003387_seg001 | VGlut-Gal4 | F | 8190.79 | 0.72 |
| VGlut-F-000514 | 9100 | DvGlutMARCM-F003408_seg001 | VGlut-Gal4 | F | 8023.19 | 0.66 |
| VGlut-F-600615 | 9144 | DvGlutMARCM-F003435_seg001 | VGlut-Gal4 | F | 6548.12 | 0.66 |
| VGlut-F-500613 | 9193 | DvGlutMARCM-F003261_seg001 | VGlut-Gal4 | F | 8074.99 | 0.67 |
| VGlut-F-700225 | 11029 | DvGlutMARCM-F002912_seg001 | VGlut-Gal4 | F | 7715.83 | 0.57 |
| VGlut-F-600307 | 12035 | DvGlutMARCM-F002551_seg005 | VGlut-Gal4 | F | 6657.34 | 0.57 |
| VGlut-F-600380 | 12237 | DvGlutMARCM-F002700_seg001 | VGlut-Gal4 | F | 8011.16 | 0.63 |
| VGlut-F-200352 | 12959 | DvGlutMARCM-F1990_seg1 | VGlut-Gal4 | F | 6762.55 | 0.65 |
| VGlut-F-300234 | 14939 | DvGlutMARCM-F1278_seg2 | VGlut-Gal4 | F | 7473.35 | 0.50 |
| VGlut-F-200040 | 15652 | DvGlutMARCM-F067_seg1 | VGlut-Gal4 | F | 7483.15 | 0.52 |
| VGlut-F-000128 | 16140 | DvGlutMARCM-F522-X2_seg2 | VGlut-Gal4 | F | 7050.52 | 0.51 |