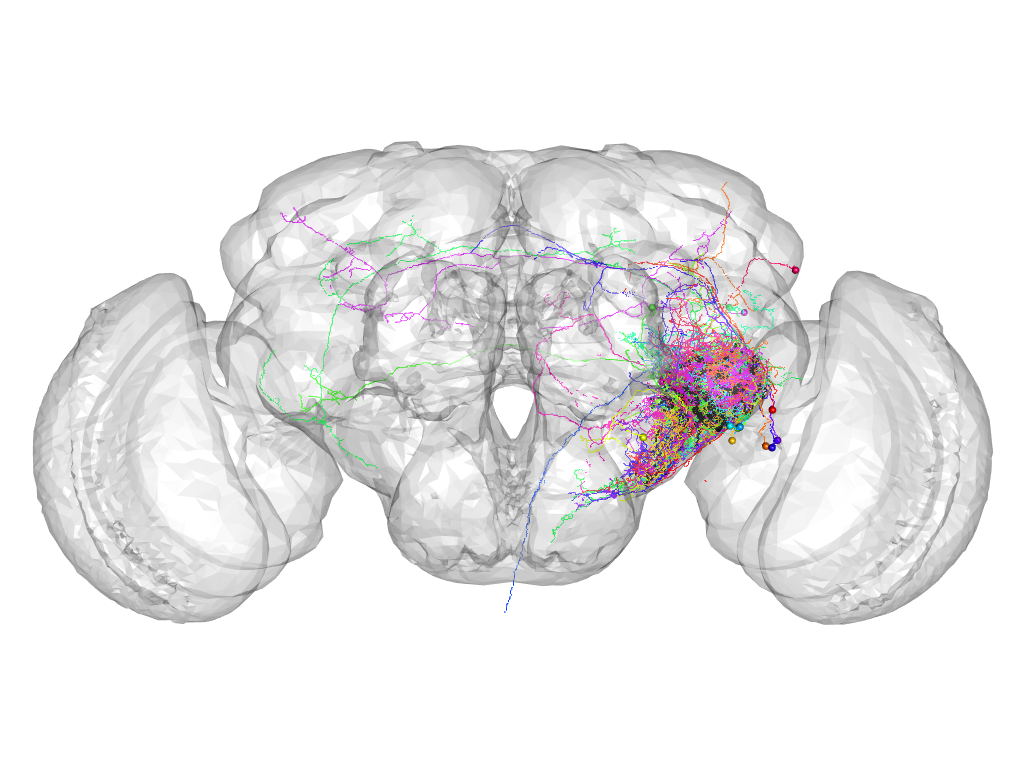
Cluster 903
Part of supercluster X
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-000273), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 200 | TH-M-000060 | X | 0.70 |
| 865 | TH-F-000025 | X | 0.70 |
| 268 | Cha-F-200340 | X | 0.64 |
| 876 | Trh-F-100023 | X | 0.63 |
| 148 | Trh-M-400144 | X | 0.57 |
Cluster composition
This cluster has 17 neurons. The exemplar of this cluster (VGlut-F-000273) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-000273 | 14365 | DvGlutMARCM-F1046_seg1 | VGlut-Gal4 | F | 71934.38 | 1.00 |
| TH-M-300046 | 3271 | THMARCM-465M_seg2 | TH-Gal4 | M | 32018.05 | 0.42 |
| Gad1-F-200149 | 4113 | GadMARCM-F000698_seg001 | Gad1-Gal4 | F | 38358.62 | 0.50 |
| Cha-F-000279 | 4152 | ChaMARCM-F001267_seg001 | Cha-Gal4 | F | 33421.98 | 0.58 |
| Cha-F-200339 | 4306 | ChaMARCM-F001370_seg002 | Cha-Gal4 | F | 41899.16 | 0.61 |
| Cha-F-300294 | 5091 | ChaMARCM-F001153_seg001 | Cha-Gal4 | F | 44268.81 | 0.61 |
| Cha-F-000251 | 5145 | ChaMARCM-F001194_seg001 | Cha-Gal4 | F | 39571.58 | 0.50 |
| Cha-F-300232 | 5501 | ChaMARCM-F000820_seg001 | Cha-Gal4 | F | 42905.44 | 0.36 |
| Cha-F-300156 | 7133 | ChaMARCM-F000601_seg001 | Cha-Gal4 | F | 42970.72 | 0.53 |
| Cha-F-600071 | 7711 | ChaMARCM-F000526_seg001 | Cha-Gal4 | F | 33644.01 | 0.61 |
| Cha-F-200106 | 7725 | ChaMARCM-F000537_seg001 | Cha-Gal4 | F | 35369.04 | 0.63 |
| Cha-F-400059 | 8373 | ChaMARCM-F000315_seg001 | Cha-Gal4 | F | 38353.85 | 0.48 |
| TH-F-400015 | 10484 | THMARCM-700F_seg | TH-Gal4 | F | 35518.28 | 0.39 |
| TH-F-000058 | 13671 | THMARCM-695F-X2_seg2 | TH-Gal4 | F | 40450.37 | 0.51 |
| VGlut-F-100084 | 14327 | DvGlutMARCM-F1010_seg3 | VGlut-Gal4 | F | 43218.20 | 0.51 |
| VGlut-F-100129 | 14906 | DvGlutMARCM-F1249_seg1 | VGlut-Gal4 | F | 47095.85 | 0.57 |
| VGlut-F-000368 | 15189 | DvGlutMARCM-F1518_seg1 | VGlut-Gal4 | F | 47050.82 | 0.61 |