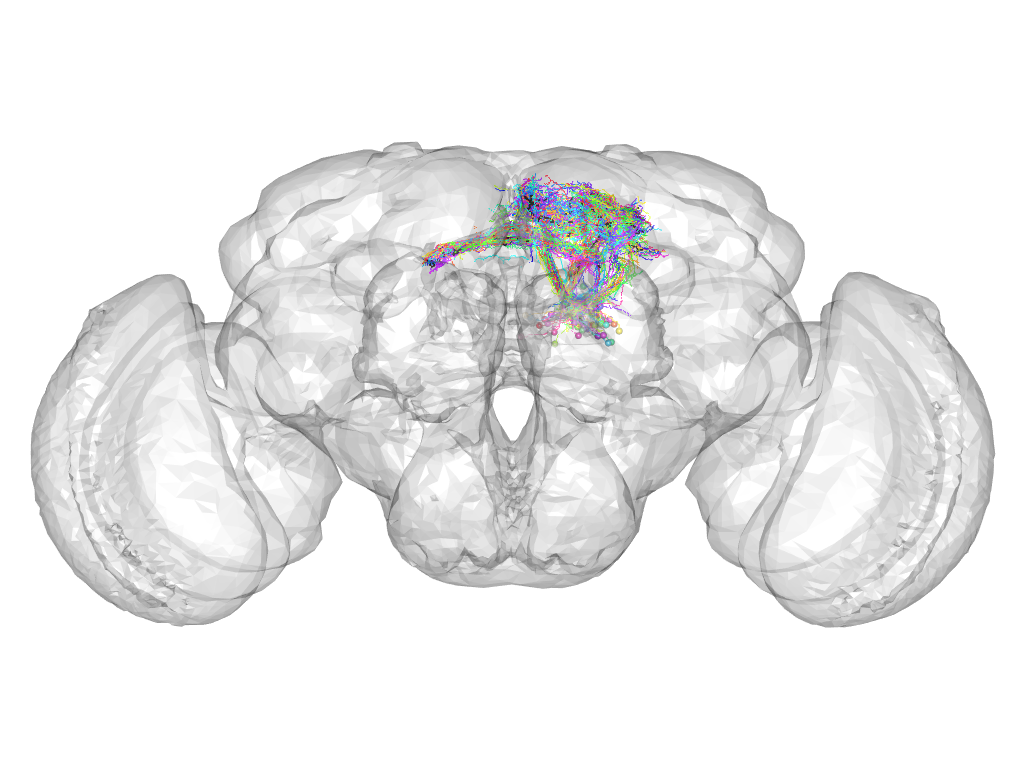
Cluster 880
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-500020), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 890 | Trh-F-300054 | XIX | 0.73 |
| 663 | Tdc2-F-200026 | XIX | 0.65 |
| 875 | Trh-F-100018 | XIX | 0.52 |
| 141 | Trh-M-400106 | XIX | 0.44 |
| 886 | Trh-F-300042 | XIX | 0.42 |
Cluster composition
This cluster has 53 neurons. The exemplar of this cluster (Trh-F-500020) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-500020 | 13895 | TPHMARCM-160F_seg1 | Trh-Gal4 | F | 16639.66 | 1.00 |
| Trh-M-400107 | 2069 | TPHMARCM-M001572_seg001 | Trh-Gal4 | M | 11991.00 | 0.73 |
| Trh-M-400118 | 2102 | TPHMARCM-M001601_seg002 | Trh-Gal4 | M | 11133.37 | 0.70 |
| Trh-M-400127 | 2138 | TPHMARCM-M001628_seg001 | Trh-Gal4 | M | 11389.61 | 0.68 |
| Trh-M-400135 | 2152 | TPHMARCM-M001640_seg001 | Trh-Gal4 | M | 11246.89 | 0.70 |
| Trh-M-400142 | 2188 | TPHMARCM-M001678_seg001 | Trh-Gal4 | M | 11651.12 | 0.72 |
| Trh-M-300073 | 2580 | TPHMARCM-851M_seg1 | Trh-Gal4 | M | 11356.76 | 0.68 |
| Trh-M-200058 | 2590 | TPHMARCM-1173M_seg1 | Trh-Gal4 | M | 10975.73 | 0.69 |
| Trh-M-300131 | 2606 | TPHMARCM-1270M_seg1 | Trh-Gal4 | M | 12121.04 | 0.71 |
| Trh-M-100077 | 2638 | TPHMARCM-M001407_seg001 | Trh-Gal4 | M | 10364.91 | 0.64 |
| Trh-M-300092 | 2785 | TPHMARCM-1009M_seg1 | Trh-Gal4 | M | 11205.98 | 0.65 |
| Trh-M-400051 | 2801 | TPHMARCM-1029M_seg1 | Trh-Gal4 | M | 9605.18 | 0.58 |
| Trh-M-300095 | 2851 | TPHMARCM-1079M_seg1 | Trh-Gal4 | M | 11835.27 | 0.72 |
| Trh-M-400086 | 2857 | TPHMARCM-1085M_seg1 | Trh-Gal4 | M | 9617.12 | 0.61 |
| Trh-M-100069 | 2902 | TPHMARCM-1141M_seg1 | Trh-Gal4 | M | 11590.27 | 0.70 |
| Trh-M-100070 | 2908 | TPHMARCM-1156M_seg1 | Trh-Gal4 | M | 10625.15 | 0.65 |
| Trh-M-300076 | 2929 | TPHMARCM-864M_seg1 | Trh-Gal4 | M | 11385.29 | 0.66 |
| Trh-M-900000 | 3450 | TPHMARCM-206M_seg1 | Trh-Gal4 | M | 9887.36 | 0.62 |
| Trh-M-200017 | 3459 | TPHMARCM-320M_seg1 | Trh-Gal4 | M | 10733.31 | 0.65 |
| Trh-M-300011 | 3468 | TPHMARCM-338M_seg1 | Trh-Gal4 | M | 11546.77 | 0.72 |
| Trh-M-300025 | 3522 | TPHMARCM-401M_seg1 | Trh-Gal4 | M | 11602.41 | 0.71 |
| Trh-M-400021 | 3550 | TPHMARCM-473M_seg1 | Trh-Gal4 | M | 10448.03 | 0.65 |
| Trh-M-100044 | 3561 | TPHMARCM-493M_seg1 | Trh-Gal4 | M | 11714.26 | 0.70 |
| Trh-M-400024 | 3586 | TPHMARCM-530M_seg1 | Trh-Gal4 | M | 11457.92 | 0.70 |
| Trh-M-200029 | 3603 | TPHMARCM-546M_seg1 | Trh-Gal4 | M | 11933.19 | 0.70 |
| Trh-M-200033 | 3695 | TPHMARCM-846M_seg1 | Trh-Gal4 | M | 11461.74 | 0.71 |
| Trh-M-200034 | 3696 | TPHMARCM-847M_seg1 | Trh-Gal4 | M | 11733.39 | 0.70 |
| Tdc2-F-400000 | 10189 | dTdc-2MARCM-F-008_seg1 | Tdc2-Gal4 | F | 11518.61 | 0.68 |
| Tdc2-F-300023 | 10303 | dTdc2MARCM-F000166_seg001 | Tdc2-Gal4 | F | 10031.34 | 0.62 |
| Tdc2-F-300025 | 10305 | dTdc2MARCM-F000168_seg001 | Tdc2-Gal4 | F | 11258.13 | 0.66 |
| Tdc2-F-300017 | 10406 | dTdc2MARCM-F000121_seg001 | Tdc2-Gal4 | F | 10998.03 | 0.67 |
| Tdc2-F-300018 | 10407 | dTdc2MARCM-F000122_seg001 | Tdc2-Gal4 | F | 11854.35 | 0.71 |
| Tdc2-F-200031 | 10410 | dTdc2MARCM-F000125_seg001 | Tdc2-Gal4 | F | 12440.48 | 0.71 |
| Tdc2-F-200032 | 10411 | dTdc2MARCM-F000126_seg001 | Tdc2-Gal4 | F | 11250.66 | 0.66 |
| Tdc2-F-400021 | 10429 | dTdc2MARCM-F000145_seg001 | Tdc2-Gal4 | F | 12248.38 | 0.74 |
| Tdc2-F-500001 | 10430 | dTdc2MARCM-F000146_seg001 | Tdc2-Gal4 | F | 11011.45 | 0.67 |
| Trh-F-200116 | 12387 | TPHMARCM-1182F_seg1 | Trh-Gal4 | F | 11483.36 | 0.69 |
| Trh-F-300149 | 12556 | TPHMARCM-1344F_seg1 | Trh-Gal4 | F | 10764.02 | 0.66 |
| Tdc2-F-000010 | 12640 | dTdc-2MARCM-F-059_seg1 | Tdc2-Gal4 | F | 11496.21 | 0.69 |
| Tdc2-F-400006 | 12659 | dTdc2MARCM-F000078_seg001 | Tdc2-Gal4 | F | 9971.95 | 0.59 |
| Tdc2-F-400007 | 12660 | dTdc2MARCM-F000079_seg001 | Tdc2-Gal4 | F | 11793.31 | 0.69 |
| Tdc2-F-200014 | 12665 | dTdc2MARCM-F000087_seg001 | Tdc2-Gal4 | F | 11775.77 | 0.70 |
| Tdc2-F-300013 | 12672 | dTdc2MARCM-F000094_seg001 | Tdc2-Gal4 | F | 12112.74 | 0.71 |
| Tdc2-F-200005 | 13194 | dTdc-2MARCM-F-025_seg1 | Tdc2-Gal4 | F | 10139.13 | 0.61 |
| Tdc2-F-300005 | 13216 | dTdc-2MARCM-F-048_seg1 | Tdc2-Gal4 | F | 11647.27 | 0.69 |
| Trh-F-200089 | 13236 | TPHMARCM-723F_seg1 | Trh-Gal4 | F | 11227.50 | 0.68 |
| Trh-F-300108 | 13240 | TPHMARCM-727F_seg1 | Trh-Gal4 | F | 11930.97 | 0.73 |
| Trh-F-200025 | 13953 | TPHMARCM-244F_seg1 | Trh-Gal4 | F | 12763.79 | 0.77 |
| Trh-F-200042 | 13981 | TPHMARCM-271F_seg1 | Trh-Gal4 | F | 11919.27 | 0.70 |
| Trh-F-200062 | 14030 | TPHMARCM-315F_seg1 | Trh-Gal4 | F | 11421.98 | 0.70 |
| Trh-F-300049 | 14076 | TPHMARCM-425F_seg1 | Trh-Gal4 | F | 11706.14 | 0.72 |
| Trh-F-300057 | 14087 | TPHMARCM-438F_seg1 | Trh-Gal4 | F | 11913.02 | 0.72 |
| Trh-F-300067 | 14122 | TPHMARCM-481F_seg2 | Trh-Gal4 | F | 12121.13 | 0.71 |