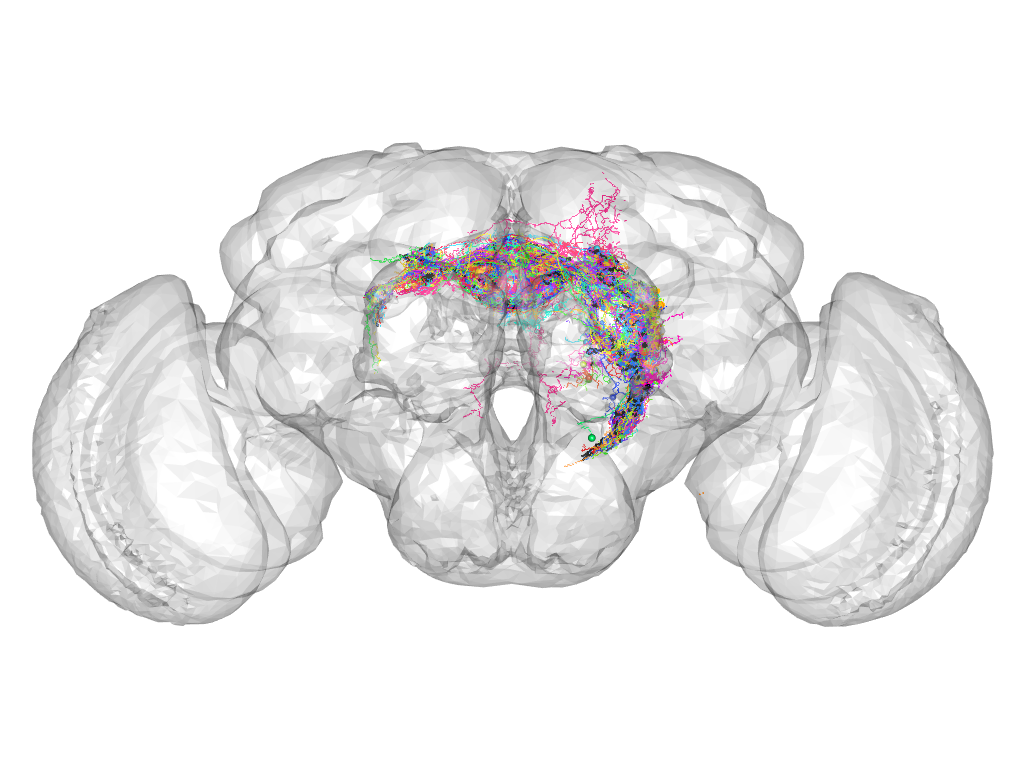
Cluster 851
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-F-100029), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1028 | VGlut-F-000067 | XIV | 0.56 |
| 964 | VGlut-F-200325 | XIV | 0.47 |
| 191 | TH-M-200018 | XIV | 0.45 |
| 192 | TH-M-300027 | XIV | 0.33 |
| 753 | VGlut-F-600178 | XIV | 0.32 |
Cluster composition
This cluster has 26 neurons. The exemplar of this cluster (TH-F-100029) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-F-100029 | 13448 | THMARCM-286F_seg1 | TH-Gal4 | F | 164061.85 | 1.00 |
| TH-M-200061 | 1720 | THMARCM-399M_seg001 | TH-Gal4 | M | 121564.79 | 0.75 |
| TH-M-100056 | 2009 | THMARCM-413M_seg1 | TH-Gal4 | M | 122023.82 | 0.75 |
| TH-M-300060 | 3073 | THMARCM-730M_seg1 | TH-Gal4 | M | 122089.53 | 0.75 |
| TH-M-100057 | 3255 | THMARCM-430M_seg1 | TH-Gal4 | M | 121498.99 | 0.75 |
| TH-M-000009 | 3268 | THMARCM-459M_seg1 | TH-Gal4 | M | 119775.17 | 0.74 |
| TH-M-100063 | 3277 | THMARCM-472M_seg1 | TH-Gal4 | M | 114611.86 | 0.73 |
| TH-M-200080 | 3281 | THMARCM-475M_seg1 | TH-Gal4 | M | 118685.01 | 0.75 |
| TH-M-300061 | 3361 | THMARCM-771M_seg1 | TH-Gal4 | M | 111511.73 | 0.72 |
| TH-M-300066 | 3369 | THMARCM-784M_seg1 | TH-Gal4 | M | 118750.20 | 0.74 |
| Trh-M-000012 | 3475 | TPHMARCM-346M_seg1 | Trh-Gal4 | M | 102685.98 | 0.66 |
| Trh-M-200019 | 3482 | TPHMARCM-354M_seg1 | Trh-Gal4 | M | 65088.42 | 0.57 |
| Cha-F-500059 | 8176 | ChaMARCM-F000194_seg001 | Cha-Gal4 | F | 110848.76 | 0.70 |
| Cha-F-000010 | 10117 | ChaMARCM-F000069_seg001 | Cha-Gal4 | F | 61849.20 | 0.48 |
| VGlut-F-600124 | 13100 | DvGlutMARCM-F2124_seg1 | VGlut-Gal4 | F | 62658.76 | 0.54 |
| TH-F-300013 | 13390 | THMARCM-158F_seg1 | TH-Gal4 | F | 123285.62 | 0.77 |
| TH-F-200023 | 13456 | THMARCM-294F_seg1 | TH-Gal4 | F | 138488.20 | 0.83 |
| TH-F-200061 | 13510 | THMARCM-385F_seg1 | TH-Gal4 | F | 121952.08 | 0.74 |
| TH-F-200073 | 13523 | THMARCM-402F_seg1 | TH-Gal4 | F | 114292.72 | 0.73 |
| TH-F-200074 | 13529 | THMARCM-412F_seg1 | TH-Gal4 | F | 114091.73 | 0.73 |
| TH-F-200075 | 13534 | THMARCM-420F_seg1 | TH-Gal4 | F | 123813.02 | 0.76 |
| TH-F-200101 | 13613 | THMARCM-560F_seg | TH-Gal4 | F | 122328.48 | 0.76 |
| TH-F-300090 | 13751 | THMARCM-788F_seg1 | TH-Gal4 | F | 116998.54 | 0.75 |
| TH-F-100001 | 13779 | THMARCM-F_017_seg1 | TH-Gal4 | F | 115206.38 | 0.71 |
| Trh-F-400005 | 13831 | TPHMARCM-052F_seg1 | Trh-Gal4 | F | 102964.96 | 0.52 |
| VGlut-F-300269 | 15160 | DvGlutMARCM-F1492_seg1 | VGlut-Gal4 | F | 62773.68 | 0.53 |