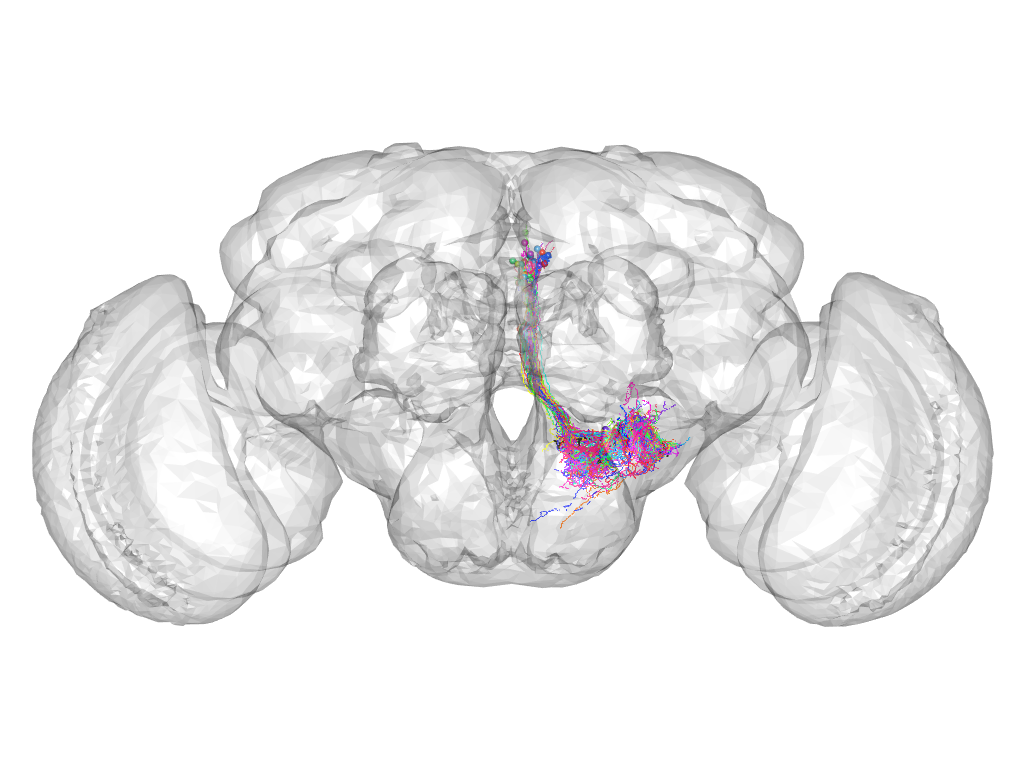
Cluster 691
Part of supercluster VIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-700120), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 690 | VGlut-F-400061 | VIII | 0.79 |
| 430 | VGlut-F-500830 | VIII | 0.76 |
| 441 | VGlut-F-600718 | VIII | 0.62 |
| 751 | VGlut-F-400648 | VIII | 0.61 |
| 685 | VGlut-F-400542 | VIII | 0.58 |
| 801 | Trh-F-600102 | VIII | 0.56 |
| 652 | Tdc2-F-200010 | VIII | 0.52 |
| 804 | Trh-F-500188 | VIII | 0.52 |
| 803 | Trh-F-700063 | VIII | 0.51 |
Cluster composition
This cluster has 59 neurons. The exemplar of this cluster (VGlut-F-700120) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-700120 | 10846 | DvGlutMARCM-F002773_seg001 | VGlut-Gal4 | F | 9999.74 | 1.00 |
| VGlut-F-500122 | 6133 | DvGlutMARCM-F870_seg1 | VGlut-Gal4 | F | 7288.51 | 0.73 |
| VGlut-F-300642 | 6590 | DvGlutMARCM-F004349_seg002 | VGlut-Gal4 | F | 7199.68 | 0.72 |
| VGlut-F-600712 | 6783 | DvGlutMARCM-F003978_seg002 | VGlut-Gal4 | F | 7365.91 | 0.71 |
| VGlut-F-500534 | 8450 | DvGlutMARCM-F003049_seg001 | VGlut-Gal4 | F | 7448.49 | 0.77 |
| VGlut-F-800132 | 8462 | DvGlutMARCM-F003060_seg001 | VGlut-Gal4 | F | 7394.01 | 0.64 |
| VGlut-F-500539 | 8480 | DvGlutMARCM-F003074_seg001 | VGlut-Gal4 | F | 7627.37 | 0.76 |
| VGlut-F-500542 | 8483 | DvGlutMARCM-F003075_seg001 | VGlut-Gal4 | F | 6936.34 | 0.73 |
| VGlut-F-500543 | 8484 | DvGlutMARCM-F003075_seg002 | VGlut-Gal4 | F | 7325.97 | 0.76 |
| VGlut-F-500576 | 8562 | DvGlutMARCM-F003129_seg001 | VGlut-Gal4 | F | 7451.23 | 0.75 |
| VGlut-F-500618 | 9198 | DvGlutMARCM-F003265_seg001 | VGlut-Gal4 | F | 7375.24 | 0.75 |
| VGlut-F-300467 | 9323 | DvGlutMARCM-F003161_seg001 | VGlut-Gal4 | F | 7131.86 | 0.70 |
| VGlut-F-300468 | 9324 | DvGlutMARCM-F003162_seg001 | VGlut-Gal4 | F | 7124.31 | 0.71 |
| VGlut-F-500598 | 9379 | DvGlutMARCM-F003210_seg001 | VGlut-Gal4 | F | 7706.34 | 0.77 |
| VGlut-F-500606 | 9387 | DvGlutMARCM-F003215_seg002 | VGlut-Gal4 | F | 7434.05 | 0.74 |
| VGlut-F-600546 | 9397 | DvGlutMARCM-F003221_seg001 | VGlut-Gal4 | F | 7271.66 | 0.75 |
| VGlut-F-300266 | 10228 | DvGlutMARCM-F1482_seg1 | VGlut-Gal4 | F | 7408.81 | 0.73 |
| VGlut-F-400210 | 10754 | DvGlutMARCM-F780_seg1 | VGlut-Gal4 | F | 7408.90 | 0.72 |
| VGlut-F-700125 | 10851 | DvGlutMARCM-F002776_seg002 | VGlut-Gal4 | F | 7196.01 | 0.72 |
| VGlut-F-700126 | 10852 | DvGlutMARCM-F002776_seg003 | VGlut-Gal4 | F | 7438.31 | 0.76 |
| VGlut-F-700155 | 10880 | DvGlutMARCM-F002788_seg001 | VGlut-Gal4 | F | 7443.29 | 0.74 |
| VGlut-F-700156 | 10881 | DvGlutMARCM-F002789_seg001 | VGlut-Gal4 | F | 7143.35 | 0.73 |
| VGlut-F-700183 | 10907 | DvGlutMARCM-F002809_seg001 | VGlut-Gal4 | F | 7496.98 | 0.72 |
| VGlut-F-600447 | 10981 | DvGlutMARCM-F002871_seg001 | VGlut-Gal4 | F | 7372.79 | 0.75 |
| VGlut-F-500494 | 11036 | DvGlutMARCM-F002916_seg002 | VGlut-Gal4 | F | 7343.78 | 0.73 |
| VGlut-F-600475 | 11119 | DvGlutMARCM-F002984_seg001 | VGlut-Gal4 | F | 7372.20 | 0.71 |
| VGlut-F-700238 | 11182 | DvGlutMARCM-F003030_seg001 | VGlut-Gal4 | F | 7420.06 | 0.75 |
| VGlut-F-600227 | 11829 | DvGlutMARCM-F002388_seg001 | VGlut-Gal4 | F | 7455.22 | 0.73 |
| VGlut-F-500528 | 11903 | DvGlutMARCM-F002441_seg001 | VGlut-Gal4 | F | 7439.79 | 0.75 |
| VGlut-F-500386 | 11937 | DvGlutMARCM-F002472_seg001 | VGlut-Gal4 | F | 7240.23 | 0.71 |
| VGlut-F-700094 | 12103 | DvGlutMARCM-F002604_seg001 | VGlut-Gal4 | F | 7457.45 | 0.76 |
| VGlut-F-600343 | 12172 | DvGlutMARCM-F002654_seg001 | VGlut-Gal4 | F | 7708.62 | 0.76 |
| VGlut-F-600373 | 12219 | DvGlutMARCM-F002686_seg002 | VGlut-Gal4 | F | 7853.25 | 0.76 |
| VGlut-F-600406 | 12274 | DvGlutMARCM-F002732_seg001 | VGlut-Gal4 | F | 7344.44 | 0.74 |
| VGlut-F-600121 | 13059 | DvGlutMARCM-F2082_seg1 | VGlut-Gal4 | F | 7575.94 | 0.76 |
| VGlut-F-600125 | 13105 | DvGlutMARCM-F2130_seg1 | VGlut-Gal4 | F | 7691.96 | 0.77 |
| VGlut-F-400295 | 14356 | DvGlutMARCM-F1039_seg1 | VGlut-Gal4 | F | 7526.11 | 0.72 |
| VGlut-F-400187 | 14454 | DvGlutMARCM-F742-x3_seg2 | VGlut-Gal4 | F | 7495.62 | 0.70 |
| VGlut-F-600036 | 14491 | DvGlutMARCM-F764_seg1 | VGlut-Gal4 | F | 7553.86 | 0.75 |
| VGlut-F-400209 | 14499 | DvGlutMARCM-F771_seg1 | VGlut-Gal4 | F | 7616.84 | 0.67 |
| VGlut-F-400213 | 14513 | DvGlutMARCM-F784_seg1 | VGlut-Gal4 | F | 7176.33 | 0.69 |
| VGlut-F-400224 | 14532 | DvGlutMARCM-F801_seg1 | VGlut-Gal4 | F | 7692.74 | 0.78 |
| VGlut-F-400247 | 14586 | DvGlutMARCM-F850_seg1 | VGlut-Gal4 | F | 7632.10 | 0.77 |
| VGlut-F-400249 | 14595 | DvGlutMARCM-F858_seg2 | VGlut-Gal4 | F | 7267.24 | 0.73 |
| VGlut-F-500128 | 14613 | DvGlutMARCM-F873_seg1 | VGlut-Gal4 | F | 7475.31 | 0.70 |
| VGlut-F-400265 | 14637 | DvGlutMARCM-F893_seg1 | VGlut-Gal4 | F | 7674.23 | 0.76 |
| VGlut-F-500154 | 15032 | DvGlutMARCM-F1372_seg1 | VGlut-Gal4 | F | 7350.94 | 0.74 |
| VGlut-F-600050 | 15053 | DvGlutMARCM-F1392_seg1 | VGlut-Gal4 | F | 6991.19 | 0.68 |
| VGlut-F-600062 | 15110 | DvGlutMARCM-F1444_seg1 | VGlut-Gal4 | F | 7352.25 | 0.72 |
| VGlut-F-600066 | 15128 | DvGlutMARCM-F1461_seg1 | VGlut-Gal4 | F | 7401.18 | 0.69 |
| VGlut-F-600068 | 15134 | DvGlutMARCM-F1467_seg1 | VGlut-Gal4 | F | 7670.50 | 0.74 |
| VGlut-F-400109 | 15487 | DvGlutMARCM-F645_seg1 | VGlut-Gal4 | F | 7747.48 | 0.73 |
| VGlut-F-400121 | 15507 | DvGlutMARCM-F658-X2_seg1 | VGlut-Gal4 | F | 7696.14 | 0.71 |
| VGlut-F-400160 | 15577 | DvGlutMARCM-F705_seg1 | VGlut-Gal4 | F | 7598.06 | 0.75 |
| VGlut-F-400030 | 15724 | DvGlutMARCM-F136-X2_seg2 | VGlut-Gal4 | F | 7151.18 | 0.61 |
| VGlut-F-500028 | 15755 | DvGlutMARCM-F164-X3_seg1 | VGlut-Gal4 | F | 6317.41 | 0.64 |
| VGlut-F-500084 | 16054 | DvGlutMARCM-F441_seg1 | VGlut-Gal4 | F | 7455.04 | 0.74 |
| VGlut-F-500098 | 16086 | DvGlutMARCM-F470_seg1 | VGlut-Gal4 | F | 7632.76 | 0.75 |
| VGlut-F-500100 | 16088 | DvGlutMARCM-F471_seg1 | VGlut-Gal4 | F | 7307.92 | 0.66 |