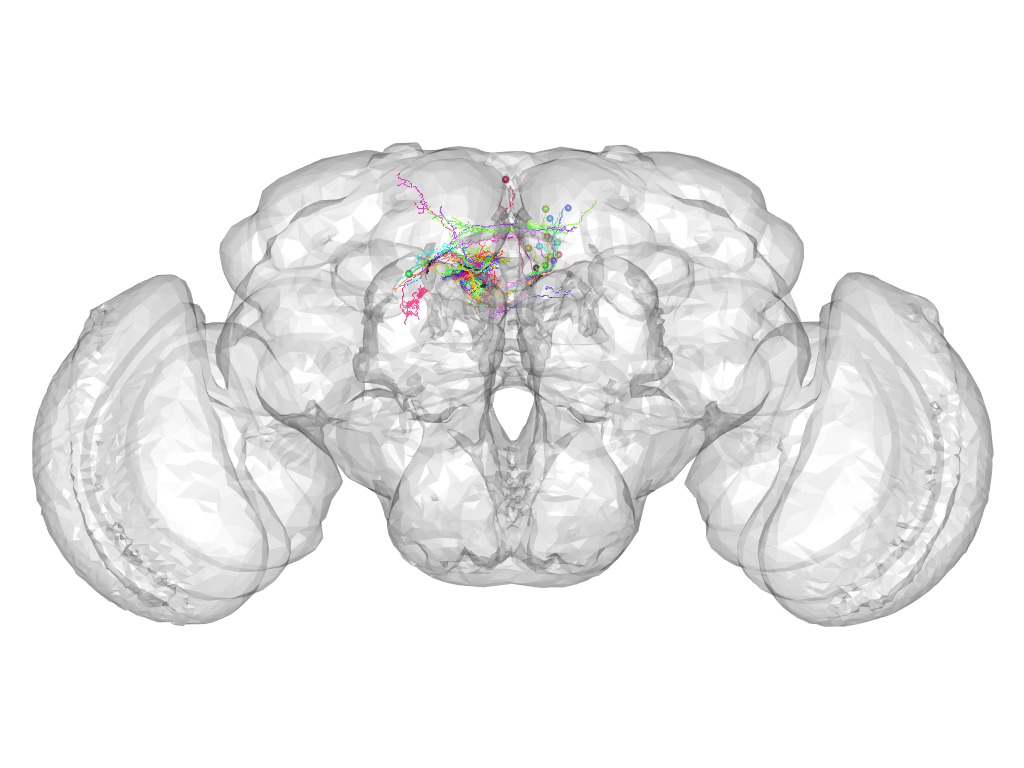
Cluster 645
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-500100), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 658 | Tdc2-F-300026 | XIV | 0.65 |
| 1028 | VGlut-F-000067 | XIV | 0.59 |
| 964 | VGlut-F-200325 | XIV | 0.57 |
| 231 | Cha-F-300338 | XIV | 0.56 |
| 416 | fru-F-600085 | XIV | 0.52 |
| 354 | Cha-F-100235 | XIV | 0.51 |
| 204 | Trh-M-100025 | XIV | 0.51 |
Cluster composition
This cluster has 17 neurons. The exemplar of this cluster (fru-F-500100) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-500100 | 10039 | FruMARCM-F000625_seg002 | fru-Gal4 | F | 6844.93 | 1.00 |
| fru-M-700149 | 78 | FruMARCM-M002277_seg001 | fru-Gal4 | M | 5242.70 | 0.72 |
| fru-M-500150 | 446 | FruMARCM-M001557_seg002 | fru-Gal4 | M | 5295.01 | 0.72 |
| fru-M-700011 | 950 | FruMARCM-M000151_seg001 | fru-Gal4 | M | 5110.65 | 0.70 |
| fru-M-500119 | 1035 | FruMARCM-M001215_seg001 | fru-Gal4 | M | 5158.71 | 0.72 |
| Gad1-F-000085 | 4092 | GadMARCM-F000681_seg001 | Gad1-Gal4 | F | 3417.20 | 0.42 |
| Gad1-F-000092 | 4104 | GadMARCM-F000692_seg001 | Gad1-Gal4 | F | 4919.97 | 0.63 |
| fru-F-500228 | 4704 | FruMARCM-F001994_seg001 | fru-Gal4 | F | 5267.05 | 0.72 |
| fru-F-500178 | 6316 | FruMARCM-F001441_seg001 | fru-Gal4 | F | 5108.11 | 0.71 |
| Cha-F-200118 | 7109 | ChaMARCM-F000583_seg001 | Cha-Gal4 | F | 4844.39 | 0.69 |
| fru-F-700065 | 7272 | FruMARCM-F000908_seg003 | fru-Gal4 | F | 5230.70 | 0.72 |
| fru-F-500113 | 7771 | FruMARCM-F000669_seg001 | fru-Gal4 | F | 5082.43 | 0.69 |
| npf-F-400006 | 9460 | npfMARCM-F000047_seg001 | npf-GAL4 | F | 4154.23 | 0.54 |
| Gad1-F-100044 | 9603 | GadMARCM-F000470_seg001 | Gad1-Gal4 | F | 4928.96 | 0.62 |
| fru-F-500031 | 9912 | FruMARCM-F000497_seg001 | fru-Gal4 | F | 5290.54 | 0.74 |
| Gad1-F-000031 | 11411 | GadMARCM-F000355_seg001 | Gad1-Gal4 | F | 4269.80 | 0.40 |
| Gad1-F-200007 | 12619 | GadMARCM-F000023_seg001 | Gad1-Gal4 | F | 4777.40 | 0.60 |