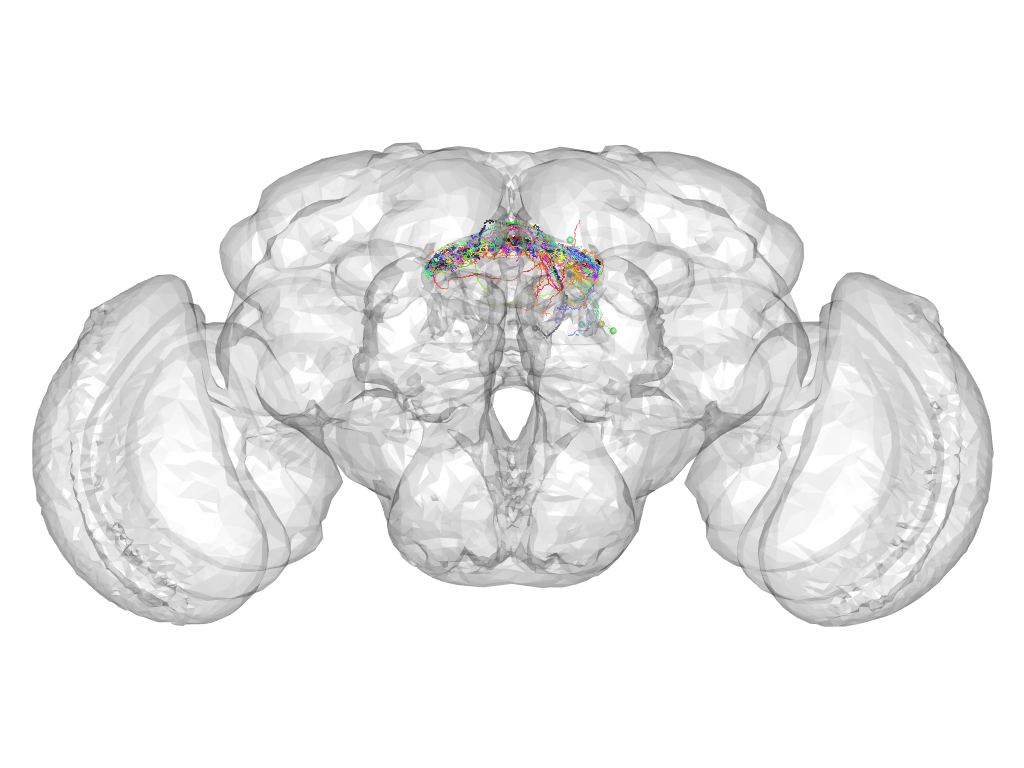
Cluster 540
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-200331), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 189 | TH-M-200006 | XIX | 0.72 |
| 83 | TH-M-300024 | XIX | 0.69 |
| 141 | Trh-M-400106 | XIX | 0.68 |
| 880 | Trh-F-500020 | XIX | 0.68 |
| 890 | Trh-F-300054 | XIX | 0.67 |
| 663 | Tdc2-F-200026 | XIX | 0.65 |
| 808 | Trh-F-300136 | XIX | 0.63 |
| 875 | Trh-F-100018 | XIX | 0.60 |
| 753 | VGlut-F-600178 | XIV | 0.58 |
| 887 | Trh-F-100046 | XIX | 0.57 |
Cluster composition
This cluster has 14 neurons. The exemplar of this cluster (VGlut-F-200331) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-200331 | 8432 | DvGlutMARCM-F1785_seg1 | VGlut-Gal4 | F | 28028.89 | 1.00 |
| fru-M-200258 | 41 | FruMARCM-M002237_seg001 | fru-Gal4 | M | 15315.32 | 0.50 |
| Trh-M-900053 | 2174 | TPHMARCM-M001664_seg001 | Trh-Gal4 | M | 20601.92 | 0.64 |
| Trh-M-400003 | 3382 | TPHMARCM-227M_seg1 | Trh-Gal4 | M | 19058.76 | 0.71 |
| Tdc2-F-100072 | 7747 | dTdc2MARCM-F000258_seg001 | Tdc2-Gal4 | F | 8656.40 | 0.38 |
| VGlut-F-200522 | 8707 | DvGlutMARCM-F003742_seg001 | VGlut-Gal4 | F | 19949.98 | 0.70 |
| Gad1-F-100054 | 9656 | GadMARCM-F000509_seg001 | Gad1-Gal4 | F | 20510.91 | 0.71 |
| Gad1-F-500092 | 9716 | GadMARCM-F000558_seg001 | Gad1-Gal4 | F | 15129.20 | 0.61 |
| VGlut-F-500519 | 11794 | DvGlutMARCM-F002360_seg001 | VGlut-Gal4 | F | 20760.56 | 0.74 |
| Gad1-F-000001 | 13181 | GadMARCM-F015_seg2 | Gad1-Gal4 | F | 10533.75 | 0.48 |
| Trh-F-300002 | 13873 | TPHMARCM-142F_seg1 | Trh-Gal4 | F | 21227.05 | 0.76 |
| Trh-F-400062 | 14199 | TPHMARCM-640F_seg1 | Trh-Gal4 | F | 20954.97 | 0.75 |
| VGlut-F-500184 | 15219 | DvGlutMARCM-F1546_seg1 | VGlut-Gal4 | F | 17916.45 | 0.66 |
| VGlut-F-500187 | 15222 | DvGlutMARCM-F1547_seg2 | VGlut-Gal4 | F | 18144.74 | 0.68 |