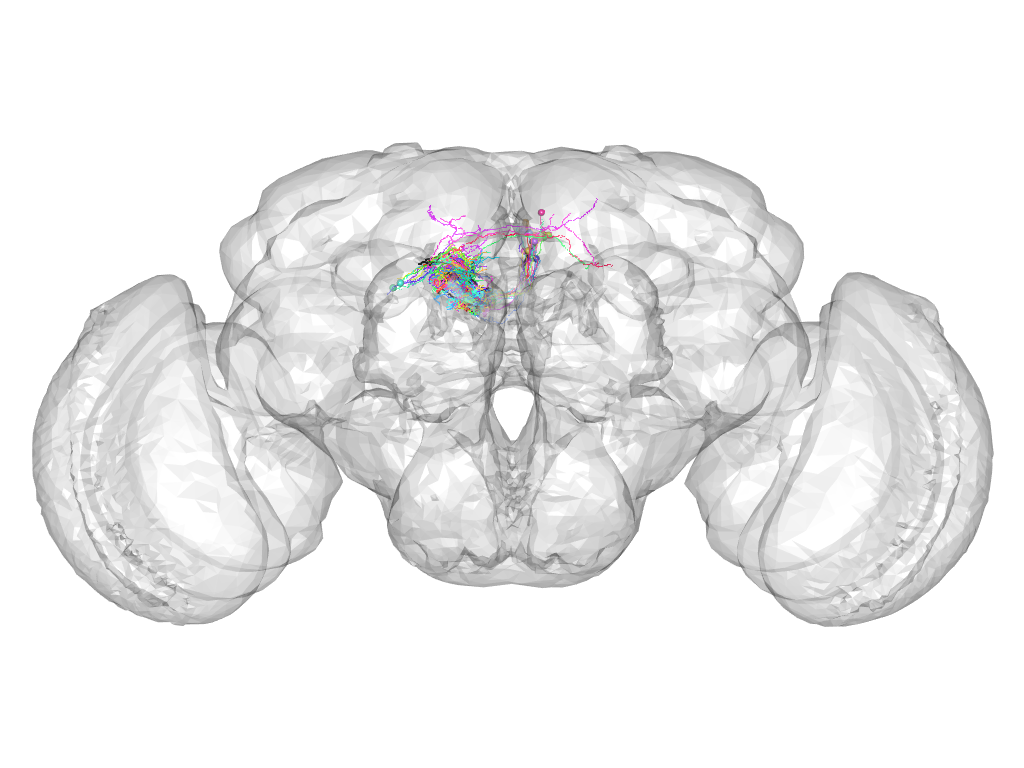
Cluster 416
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-600085), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 964 | VGlut-F-200325 | XIV | 0.62 |
| 1028 | VGlut-F-000067 | XIV | 0.59 |
| 658 | Tdc2-F-300026 | XIV | 0.54 |
| 659 | Tdc2-F-300029 | XIV | 0.51 |
| 204 | Trh-M-100025 | XIV | 0.49 |
Cluster composition
This cluster has 22 neurons. The exemplar of this cluster (fru-F-600085) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-600085 | 6390 | FruMARCM-F001526_seg001 | fru-Gal4 | F | 8803.87 | 1.00 |
| fru-M-600085 | 695 | FruMARCM-M001898_seg001 | fru-Gal4 | M | 6719.60 | 0.75 |
| fru-M-700071 | 1435 | FruMARCM-M000960_seg001 | fru-Gal4 | M | 6431.67 | 0.76 |
| fru-M-200100 | 1465 | FruMARCM-M001025_seg001 | fru-Gal4 | M | 6440.15 | 0.72 |
| fru-M-400017 | 1801 | FruMARCM-M000327_seg001 | fru-Gal4 | M | 6548.80 | 0.73 |
| fru-M-900019 | 2438 | FruMARCM-M000154_seg002 | fru-Gal4 | M | 6538.85 | 0.73 |
| Trh-M-500153 | 2545 | TPHMARCM-M001524_seg001 | Trh-Gal4 | M | 6463.05 | 0.73 |
| Trh-M-400050 | 2799 | TPHMARCM-1027M_seg1 | Trh-Gal4 | M | 6254.08 | 0.74 |
| Cha-F-600166 | 4199 | ChaMARCM-F001296_seg001 | Cha-Gal4 | F | 6469.26 | 0.70 |
| fru-F-500219 | 4661 | FruMARCM-F001824_seg001 | fru-Gal4 | F | 4904.64 | 0.52 |
| fru-F-600061 | 6176 | FruMARCM-F001110_seg003 | fru-Gal4 | F | 6371.90 | 0.70 |
| fru-F-600065 | 6226 | FruMARCM-F001160_seg001 | fru-Gal4 | F | 6781.87 | 0.74 |
| fru-F-500171 | 6267 | FruMARCM-F001302_seg001 | fru-Gal4 | F | 6555.27 | 0.73 |
| fru-F-500186 | 6324 | FruMARCM-F001446_seg001 | fru-Gal4 | F | 5915.69 | 0.65 |
| Cha-F-000096 | 7099 | ChaMARCM-F000575_seg001 | Cha-Gal4 | F | 4321.26 | 0.53 |
| fru-F-500141 | 7825 | FruMARCM-F000719_seg001 | fru-Gal4 | F | 6531.49 | 0.74 |
| fru-F-500017 | 9849 | FruMARCM-F000433_seg001 | fru-Gal4 | F | 6406.89 | 0.71 |
| fru-F-400072 | 9923 | FruMARCM-F000504_seg001 | fru-Gal4 | F | 6656.00 | 0.74 |
| Cha-F-500029 | 10172 | ChaMARCM-F000102_seg001 | Cha-Gal4 | F | 5615.46 | 0.48 |
| Gad1-F-200089 | 11335 | GadMARCM-F000303_seg001 | Gad1-Gal4 | F | 6619.56 | 0.64 |
| fru-F-700010 | 11454 | FruMARCM-F000070_seg002 | fru-Gal4 | F | 6285.83 | 0.70 |
| Cha-F-500000 | 11577 | ChaMARCM-F000014_seg001 | Cha-Gal4 | F | 6087.03 | 0.60 |