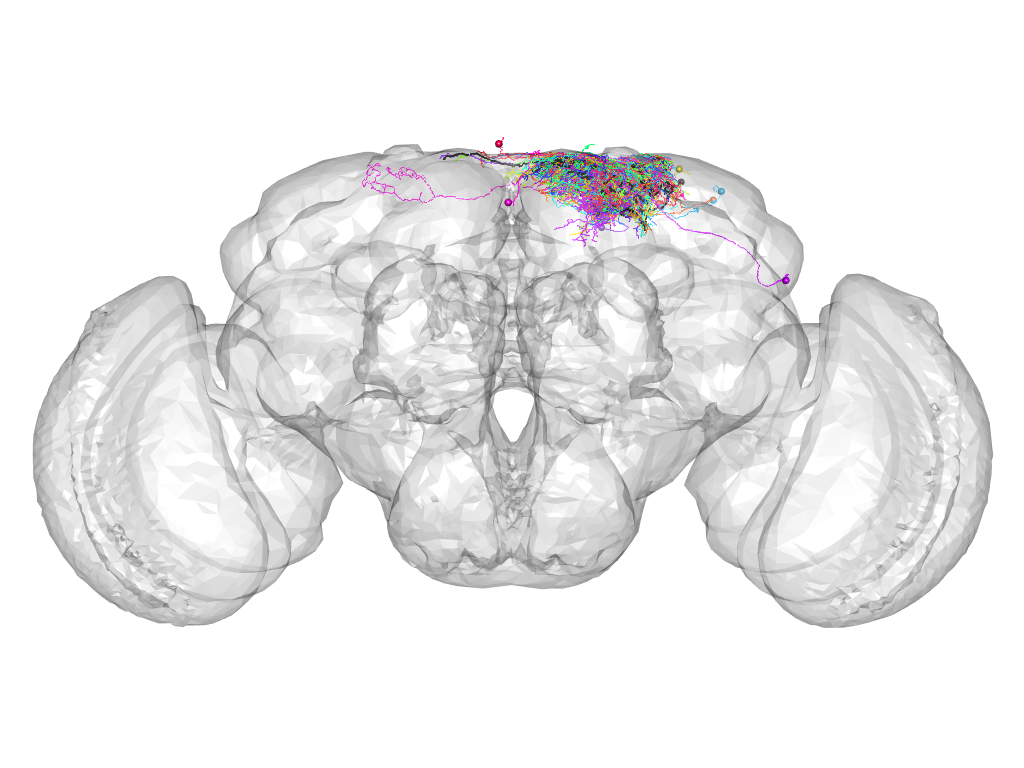
Cluster 39
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-000219), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 881 | Trh-F-200023 | XIX | 0.75 |
| 1003 | VGlut-F-000024 | XIX | 0.72 |
| 193 | TH-M-100039 | XIX | 0.66 |
| 992 | VGlut-F-200024 | XIX | 0.60 |
| 40 | fru-M-000123 | XIX | 0.58 |
| 850 | TH-F-100027 | XIX | 0.57 |
| 1047 | VGlut-F-000158 | XIX | 0.54 |
| 83 | TH-M-300024 | XIX | 0.51 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (fru-M-000219) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-000219 | 409 | FruMARCM-M002649_seg001 | fru-Gal4 | M | 36229.14 | 1.00 |
| fru-M-200310 | 313 | FruMARCM-M002538_seg001 | fru-Gal4 | M | 26789.84 | 0.74 |
| fru-M-000215 | 384 | FruMARCM-M002618_seg002 | fru-Gal4 | M | 26667.34 | 0.74 |
| fru-M-100236 | 404 | FruMARCM-M002644_seg001 | fru-Gal4 | M | 26655.42 | 0.73 |
| fru-M-100154 | 478 | FruMARCM-M001581_seg001 | fru-Gal4 | M | 25356.51 | 0.72 |
| fru-M-200214 | 493 | FruMARCM-M001590_seg002 | fru-Gal4 | M | 26044.65 | 0.73 |
| fru-M-000096 | 565 | FruMARCM-M001739_seg002 | fru-Gal4 | M | 26796.73 | 0.74 |
| fru-M-300229 | 748 | FruMARCM-M001963_seg001 | fru-Gal4 | M | 26013.37 | 0.74 |
| fru-M-300123 | 970 | FruMARCM-M001145_seg002 | fru-Gal4 | M | 25493.01 | 0.72 |
| fru-M-900030 | 1140 | FruMARCM-M001333_seg001 | fru-Gal4 | M | 24215.10 | 0.70 |
| fru-M-400141 | 1238 | FruMARCM-M001420_seg001 | fru-Gal4 | M | 25719.39 | 0.73 |
| fru-M-300158 | 1252 | FruMARCM-M001437_seg004 | fru-Gal4 | M | 26206.61 | 0.74 |
| fru-M-700065 | 1399 | FruMARCM-M000900_seg001 | fru-Gal4 | M | 26974.74 | 0.75 |
| fru-M-500051 | 1656 | FruMARCM-M000805_seg001 | fru-Gal4 | M | 25922.32 | 0.73 |
| fru-M-100046 | 1916 | FruMARCM-M000473_seg002 | fru-Gal4 | M | 26244.00 | 0.73 |
| fru-M-500034 | 1957 | FruMARCM-M000599_seg002 | fru-Gal4 | M | 25255.63 | 0.73 |
| fru-F-500281 | 3822 | FruMARCM-F002633_seg002 | fru-Gal4 | F | 22872.28 | 0.65 |
| Cha-F-700180 | 4449 | ChaMARCM-F001491_seg001 | Cha-Gal4 | F | 24639.87 | 0.64 |
| VGlut-F-300397 | 6110 | DvGlutMARCM-F2181_seg2 | VGlut-Gal4 | F | 22641.46 | 0.58 |
| fru-F-200079 | 6237 | FruMARCM-F001217_seg001 | fru-Gal4 | F | 25754.39 | 0.73 |
| Gad1-F-200042 | 10680 | GadMARCM-F000184_seg002 | Gad1-Gal4 | F | 19133.15 | 0.57 |