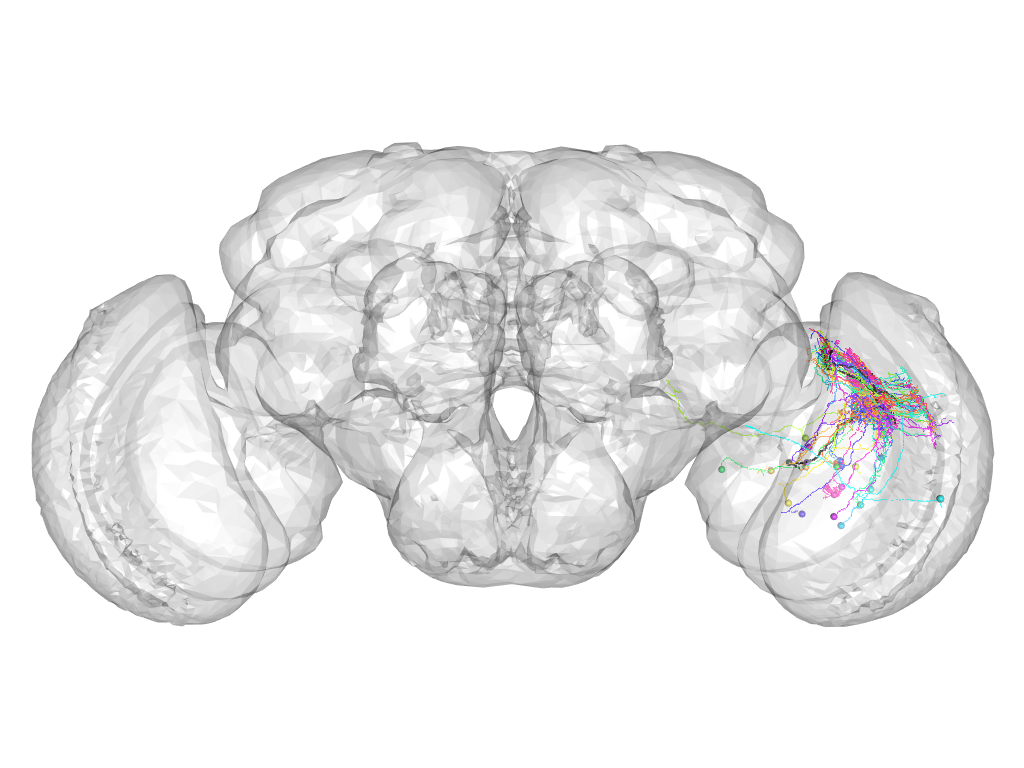
Cluster 255
Part of supercluster I
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-000282), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 935 | VGlut-F-100111 | V | 0.53 |
| 633 | fru-F-100023 | II | 0.53 |
| 133 | fru-M-700043 | I | 0.52 |
| 975 | VGlut-F-400125 | I | 0.31 |
| 327 | Cha-F-100171 | I | 0.31 |
Cluster composition
This cluster has 27 neurons. The exemplar of this cluster (Cha-F-000282) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-000282 | 4156 | ChaMARCM-F001270_seg001 | Cha-Gal4 | F | 16366.32 | 1.00 |
| Cha-F-500112 | 7707 | ChaMARCM-F000523_seg003 | Cha-Gal4 | F | 4417.48 | 0.43 |
| VGlut-F-900103 | 7932 | DvGlutMARCM-F003770_seg001 | VGlut-Gal4 | F | 8763.91 | 0.56 |
| VGlut-F-000596 | 7953 | DvGlutMARCM-F003783_seg002 | VGlut-Gal4 | F | 8948.00 | 0.57 |
| VGlut-F-000599 | 7958 | DvGlutMARCM-F003788_seg001 | VGlut-Gal4 | F | 8222.26 | 0.51 |
| VGlut-F-600660 | 8975 | DvGlutMARCM-F003535_seg001 | VGlut-Gal4 | F | 9559.30 | 0.58 |
| VGlut-F-700329 | 9293 | DvGlutMARCM-F003341_seg001 | VGlut-Gal4 | F | 8942.15 | 0.57 |
| Gad1-F-200099 | 9484 | GadMARCM-F000378_seg001 | Gad1-Gal4 | F | 8455.14 | 0.37 |
| Gad1-F-300133 | 9712 | GadMARCM-F000554_seg001 | Gad1-Gal4 | F | 4505.01 | 0.37 |
| Gad1-F-800005 | 10603 | GadMARCM-F000113_seg002 | Gad1-Gal4 | F | 7349.57 | 0.56 |
| VGlut-F-700236 | 11177 | DvGlutMARCM-F003024_seg001 | VGlut-Gal4 | F | 11063.42 | 0.65 |
| VGlut-F-400565 | 12021 | DvGlutMARCM-F002540_seg003 | VGlut-Gal4 | F | 11577.77 | 0.67 |
| VGlut-F-600308 | 12036 | DvGlutMARCM-F002551_seg006 | VGlut-Gal4 | F | 9157.52 | 0.50 |
| VGlut-F-200230 | 12370 | DvGlutMARCM-F1173_seg3 | VGlut-Gal4 | F | 8955.38 | 0.51 |
| VGlut-F-000406 | 12773 | DvGlutMARCM-F1828_seg1 | VGlut-Gal4 | F | 8298.88 | 0.32 |
| VGlut-F-100200 | 12798 | DvGlutMARCM-F1844_seg1 | VGlut-Gal4 | F | 9156.72 | 0.49 |
| VGlut-F-500296 | 12835 | DvGlutMARCM-F1881_seg2 | VGlut-Gal4 | F | 8724.20 | 0.58 |
| VGlut-F-400473 | 12931 | DvGlutMARCM-F1969_seg1 | VGlut-Gal4 | F | 10828.40 | 0.69 |
| VGlut-F-400234 | 14548 | DvGlutMARCM-F816_seg1 | VGlut-Gal4 | F | 10362.56 | 0.63 |
| VGlut-F-400319 | 14799 | DvGlutMARCM-F1149_seg1 | VGlut-Gal4 | F | 9113.87 | 0.53 |
| VGlut-F-400357 | 14964 | DvGlutMARCM-F1302_seg1 | VGlut-Gal4 | F | 11451.87 | 0.70 |
| VGlut-F-400364 | 15012 | DvGlutMARCM-F1352_seg1 | VGlut-Gal4 | F | 10748.40 | 0.60 |
| VGlut-F-300300 | 15254 | DvGlutMARCM-F1574_seg1 | VGlut-Gal4 | F | 9189.17 | 0.49 |
| VGlut-F-400162 | 15579 | DvGlutMARCM-F706-X2_seg2 | VGlut-Gal4 | F | 10821.35 | 0.60 |
| VGlut-F-000059 | 15979 | DvGlutMARCM-F370_seg1 | VGlut-Gal4 | F | 10754.57 | 0.63 |
| VGlut-F-500089 | 16060 | DvGlutMARCM-F446-X2_seg2 | VGlut-Gal4 | F | 9493.33 | 0.64 |
| VGlut-F-300091 | 16204 | DvGlutMARCM-F584_seg1 | VGlut-Gal4 | F | 9135.37 | 0.53 |