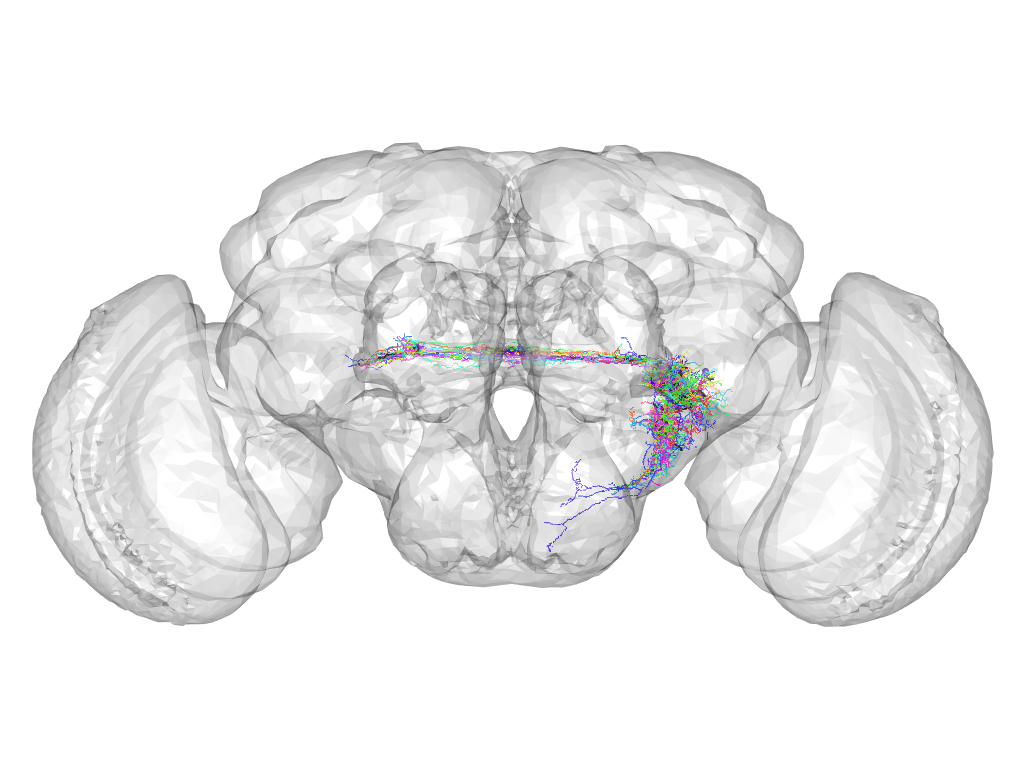
Cluster 183
Part of supercluster VII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-500074), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 148 | Trh-M-400144 | X | 0.62 |
| 200 | TH-M-000060 | X | 0.44 |
| 865 | TH-F-000025 | X | 0.39 |
| 532 | Cha-F-300093 | X | 0.34 |
| 177 | Trh-M-300109 | VIII | 0.30 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (Trh-M-500074) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-500074 | 3014 | TPHMARCM-966M_seg1 | Trh-Gal4 | M | 11207.00 | 1.00 |
| Trh-M-300088 | 1990 | TPHMARCM-0926M_seg001 | Trh-Gal4 | M | 7960.02 | 0.73 |
| Trh-M-900054 | 2175 | TPHMARCM-M001664_seg002 | Trh-Gal4 | M | 8410.84 | 0.73 |
| Trh-M-400145 | 2268 | TPHMARCM-M001756_seg001 | Trh-Gal4 | M | 8565.82 | 0.77 |
| Trh-M-500086 | 2762 | TPHMARCM-1068M_seg1 | Trh-Gal4 | M | 8499.38 | 0.77 |
| Trh-M-500083 | 2823 | TPHMARCM-1052M_seg1 | Trh-Gal4 | M | 8031.62 | 0.74 |
| Trh-M-400066 | 2834 | TPHMARCM-1061M_seg1 | Trh-Gal4 | M | 7486.19 | 0.65 |
| Trh-M-300117 | 2910 | TPHMARCM-1157M_seg1 | Trh-Gal4 | M | 7768.26 | 0.71 |
| Trh-M-300083 | 2968 | TPHMARCM-909M_seg1 | Trh-Gal4 | M | 8270.29 | 0.75 |
| Trh-M-500073 | 3011 | TPHMARCM-963M_seg1 | Trh-Gal4 | M | 7144.71 | 0.61 |
| Trh-M-100038 | 3542 | TPHMARCM-452M_seg1 | Trh-Gal4 | M | 7688.97 | 0.70 |
| Trh-M-400022 | 3573 | TPHMARCM-503M_seg1 | Trh-Gal4 | M | 8383.58 | 0.75 |
| Trh-M-300069 | 3685 | TPHMARCM-798M_seg2 | Trh-Gal4 | M | 7183.51 | 0.61 |
| VGlut-F-400809 | 6729 | DvGlutMARCM-F003938_seg001 | VGlut-Gal4 | F | 7800.78 | 0.71 |
| VGlut-F-500731 | 7422 | DvGlutMARCM-F003877_seg001 | VGlut-Gal4 | F | 5395.92 | 0.44 |
| Tdc2-F-500003 | 10299 | dTdc2MARCM-F000162_seg002 | Tdc2-Gal4 | F | 8071.99 | 0.59 |
| VGlut-F-600457 | 10993 | DvGlutMARCM-F002883_seg001 | VGlut-Gal4 | F | 8687.50 | 0.77 |
| VGlut-F-500492 | 11032 | DvGlutMARCM-F002913_seg001 | VGlut-Gal4 | F | 8156.66 | 0.75 |
| Trh-F-500091 | 13280 | TPHMARCM-767F_seg1 | Trh-Gal4 | F | 8299.69 | 0.72 |
| Trh-F-400027 | 13941 | TPHMARCM-233F-repeated0811-04_seg1 | Trh-Gal4 | F | 8034.43 | 0.74 |
| Trh-F-300092 | 14188 | TPHMARCM-630F_seg1 | Trh-Gal4 | F | 8639.09 | 0.78 |