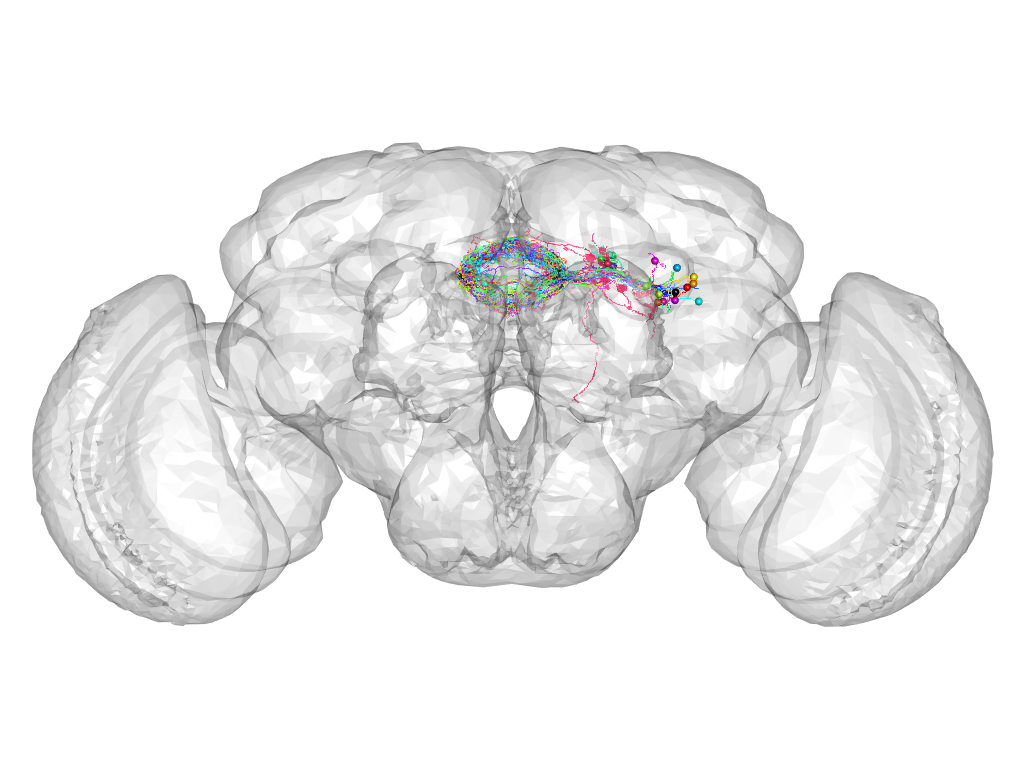
Cluster 164
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (5-HT1B-M-400020), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 851 | TH-F-100029 | XIV | 0.75 |
| 964 | VGlut-F-200325 | XIV | 0.69 |
| 1028 | VGlut-F-000067 | XIV | 0.56 |
| 149 | Trh-M-000082 | XIV | 0.51 |
| 449 | VGlut-F-600742 | XIV | 0.46 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (5-HT1B-M-400020) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| 5-HT1B-M-400020 | 2537 | 5HT1bMARCM-M000076_seg001 | 5-HT1B-Gal4 | M | 7254.94 | 1.00 |
| 5-HT1B-M-500000 | 2487 | 5HT1bMARCM-M000033_seg001 | 5-HT1B-Gal4 | M | 5644.25 | 0.77 |
| 5-HT1B-M-400001 | 2497 | 5HT1bMARCM-M000043_seg001 | 5-HT1B-Gal4 | M | 5653.43 | 0.77 |
| 5-HT1B-M-300001 | 2511 | 5HT1bMARCM-M000054_seg001 | 5-HT1B-Gal4 | M | 5852.14 | 0.76 |
| 5-HT1B-M-400013 | 2521 | 5HT1bMARCM-M000062_seg001 | 5-HT1B-Gal4 | M | 5790.92 | 0.78 |
| 5-HT1B-M-500007 | 2525 | 5HT1bMARCM-M000066_seg001 | 5-HT1B-Gal4 | M | 5655.72 | 0.78 |
| 5-HT1B-M-400018 | 2528 | 5HT1bMARCM-M000069_seg001 | 5-HT1B-Gal4 | M | 5345.65 | 0.74 |
| Gad1-F-000086 | 3937 | GadMARCM-F000682_seg001 | Gad1-Gal4 | F | 5580.30 | 0.75 |
| Gad1-F-100064 | 3942 | GadMARCM-F000572_seg001 | Gad1-Gal4 | F | 5490.19 | 0.73 |
| Gad1-F-000080 | 4080 | GadMARCM-F000670_seg001 | Gad1-Gal4 | F | 5688.27 | 0.76 |
| Gad1-F-000089 | 4101 | GadMARCM-F000690_seg001 | Gad1-Gal4 | F | 5652.69 | 0.76 |
| VGlut-F-500874 | 5884 | DvGlutMARCM-F004566_seg001 | VGlut-Gal4 | F | 5509.46 | 0.72 |
| VGlut-F-200494 | 8742 | DvGlutMARCM-F003568_seg002 | VGlut-Gal4 | F | 4904.89 | 0.64 |
| VGlut-F-200478 | 8970 | DvGlutMARCM-F003530_seg001 | VGlut-Gal4 | F | 5414.40 | 0.74 |
| Gad1-F-000050 | 9566 | GadMARCM-F000440_seg001 | Gad1-Gal4 | F | 5621.62 | 0.76 |
| Gad1-F-700047 | 9707 | GadMARCM-F000549_seg002 | Gad1-Gal4 | F | 5659.11 | 0.76 |
| Gad1-F-400007 | 10539 | GadMARCM-F000070_seg001 | Gad1-Gal4 | F | 5084.62 | 0.68 |
| 5-HT1B-F-500022 | 11203 | 5HT1bMARCM-F000032_seg001 | 5-HT1B-Gal4 | F | 5624.38 | 0.77 |
| 5-HT1B-F-500002 | 11607 | 5HT1bMARCM-F000004_seg001 | 5-HT1B-Gal4 | F | 5781.90 | 0.77 |
| 5-HT1B-F-400002 | 11627 | 5HT1bMARCM-F000020_seg001 | 5-HT1B-Gal4 | F | 5806.29 | 0.79 |
| VGlut-F-000065 | 15991 | DvGlutMARCM-F380_seg1 | VGlut-Gal4 | F | 4780.31 | 0.53 |